Projects
Below are brief summaries of various academic projects that I have worked on through the years. For software-related side projects and coding examples, please check out my GitHub profile.
Research
Constraints on the dynamics of neural population activity
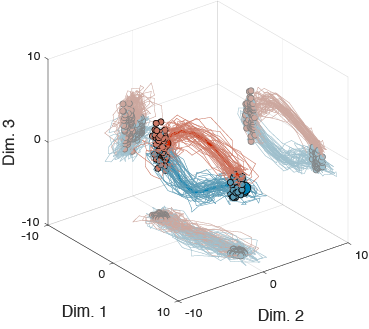
Though chiefly envisioned as a tool for the restoration of function following paralysis, brain-computer interfaces (BCIs) can serve as a powerful tool for scientific discovery. In a BCI, the mapping, which defines the relationship between the recorded neural activity and the effector (e.g., a computer cursor), is known. Therefore, by modifying the BCI mapping, we are able to “request” neural activity with specific properties. My current research focuses on using BCIs to probe the dynamics of neural population activity in motor cortex. This will allow us to better understand how the cortex carries out the computations required to guide complex behavior.
Additional information
- Degenhart, A.D. et al. Constraints on the time course of neural population activity. Cosyne Abstracts, 2020.
- Degenhart, A.D. et al. Constraints on the time course of neural population activity. Society for Neuroscience Annual Meeting 2019.
A stabilized brain-computer interface based on neural manifold alignment
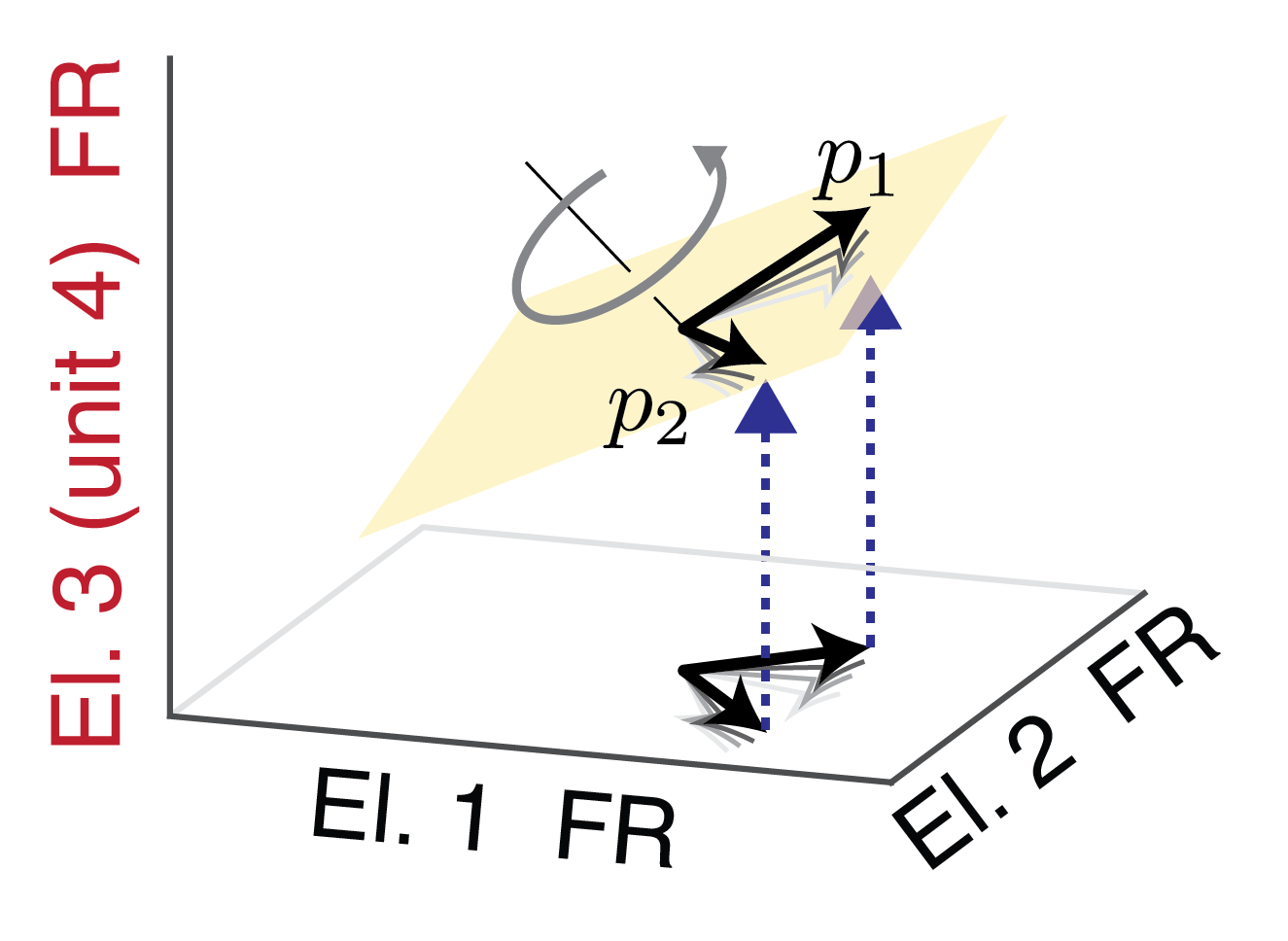
Recent demonstrations have provided compelling evidence of the ability of BCIs to provide functional restoration for individuals with paralysis. However, a major barrier to the clinical adoption of BCI systems are neural recording instabilities, which can quickly lead to degradations in device performance. To overcome this shortcoming, I, in collaboration with colleagues at Carnegie Mellon University and the University of Pittsburgh, have developed a stabilized BCI which utilizes the structure in neural population activity to improve BCI stability.
Additional information
- Degenhart, A.D.*, Bishop, W.E.*, Oby E.R., Tyler-Kabara E.C., Chase S.M.^, Batista A.P.^, Yu B.M.^ Stabilization of a brain–computer interface via the alignment of low-dimensional spaces of neural activity. Nature Biomedical Engineering (2020). (*,^: denotes equal contributions) https://doi.org/10.1038/s41551-020-0542-9
- Stabilizing brain-computer interfaces (CMU press release)
- Mind Over Body: Improving Brain-Computer Interfaces (Pitt press release)
Investigating learning using brain-computer interfaces
Similar to how BCIs can be used to probe constraints on the dynamics of neural population activity, we can leverage our ability to modify the mapping between neural activity and the BCI effector to study learning in the brain. This work, led by Emily Oby, investigated whether subjects were able to generate new neural activity patterns to overcome perturbations applied to the BCI mapping. We found that this was indeed possible, but required long-term practice across multiple days. This work provides insight into the mechanisms underlying skill learning in the brain, and has implications for how clinical BCI control could be improved through long-term training.
Additional information
- Oby, E.R., Golub, M.D., Hennig, J.A., Degenhart, A.D., Tyler-Kabara, E.C., Yu, B.M., Chase, S.M., Batista, A.B. New neural activity patterns emerge with long-term learning. Proceedings of the National Academy of Sciences (PNAS) (2019). https://doi.org/10.1073/pnas.1820296116
- Pitt and CMU Researchers Discover How the Brain Changes When Mastering a New Skill (Pitt press release)
Three-dimensional BCI control by individuals with upper-limb paralysis using electrocorticography
Electrocorticography (ECoG) has gained prominence as a potential source of control signals for BCIs, owing in part to its stability, relatively high spatial and temporal resolution (as compared to EEG), and minimal signal processing requirements. As a graduate student, I assessed the ability of individuals with upper-limb paralysis to control a three-dimensional ECoG-based BCI.
Additional information
- Degenhart, A.D., et al. Remapping cortical modulation for electrocorticographic brain–computer interfaces: a somatotopy-based approach in individuals with upper-limb paralysis. Journal of Neural Engineering (2018). https://doi.org/10.1088/1741-2552/aa9bfb
- Wang, W., Collinger, J.L., Degenhart, A.D., Tyler-Kabara, E.C., Schwartz, A.B., Moran, D.W., et al. An Electrocorticographic Brain Interface in an Individual with Tetraplegia. PLoS ONE, e55344 (2013). https://doi.org/10.1371/journal.pone.0055344
- Brain linked to robotic hand; success hailed (Pittsburgh Post-Gazette)
- 10 World-Changing Innovators for 2012 (Popular mechanics)
- Paralyzed Man Moves Robotic Arm With His Thoughts - UPMC (YouTube)
Methods and software
A modular software framework for brain-computer interface research
As part of my graduate research, I led the development of a modular software framework to enable brain-computer interface research. Our framework allowed for real-time signal acquisition, processing, decoding, control, stimulus presentation, and data saving. A key requirement of this framework was that it was highly customizable, providing control and visualization of the state of the system while maintaining high performance. This software package enabled a number of scientific studies, including our demonstration of three-dimensional BCI control by individuals with upper-limb paralysis.
Additional information
- Degenhart, A.D., Kelly, J.W., Ashmore, R.C., Collinger, J.L., Tyler-Kabara, E.C., Weber, D.J., & Wang, W. Craniux: A LabVIEW-Based Modular Software Framework for Brain-Machine Interface Research. Computational Intelligence and Neuroscience (2011). https://doi.org/10.1155/2011/363565
Improving decoding of information from Electrocorticographic recordings using Empirical Bayes
The primary focus of my graduate research was the extraction (or decoding) of information from neural recordings. A key challenge for decoding information from electrocorticograpy (ECoG) is that signal features can be extremely high-dimensional yet highly correlated. To overcome this challenge, I developed a novel decoding algorithm which uses empirical Bayes in conjunction with weight-space covariance priors to find spectrally and spatially-constrained decoding weight solutions. Using both simulated data and ECoG recordings from individuals with upper-limb paralysis, I found that by imposing prior distributions over decoding weights which encourage spectral and spatial correlation, decoding performance can be achieved which surpasses that of standard approaches. Additionally, the performance improvements of this method were found to be greatest in data-limited regimes, making it particularly appropriate for ECoG BCI systems.